Social Network Sensors for Early Detection of Contagious Outbreaks
- Authors: Nicholas A. Christakis, James H. Fowler
- Publication, Year: PLoS One, 2010
- Link to Article
Social Network Sensors for Early Detection of Contagious OutbreaksSummaryIntroductionCentral Nodes versus Random PopulationUsing Friends as Proxy for Central NodesEvaluating the Effectiveness of this MethodImportant ClarificationResultsFriend Sample versus Random Sample Early PredictionNetwork StructureCouple of Discussion Points
Summary
The authors propose a network science method to predict epidemic spreading much earlier than other methods
The method basically works like this:
Take a random sample of individuals within a population/network of interest
Ask them to self—report friends
Build a network from these reported friends and track them as a proxy for more centrally located nodes
- This approach is based on the friendship-paradox which shows us that our friends tend to have more friends than we do, on average
- Thus, these individuals should display a higher k-core and degree (on avg.) and be exposed to a spreading phenomenon much sooner than the random sample
- Essentially, it is a trick to find solid proxies for the core of a network, which we know tends to be crucially involved in spreading phenomena
Using real—world flu—spreading data (Harvard students), they illustrate that this method predicts the growth of flu infection within this network significantly sooner than the random sample (a proxy for the general network/population)
Introduction
During a contagious outbreak, individuals at the center of a network are likely to be infected sooner than random members of the population
Hence, collecting info. from a sample of central individuals within a network could be used to detect contagious outbreaks before they happen in the population-at-large
A contagion that infects individuals stochastically will tend to reach central nodes more quickly than more peripheral nodes
- This is because central nodes (defined in a few different ways) are a smaller number of steps (degrees of separation) from the average node within that network
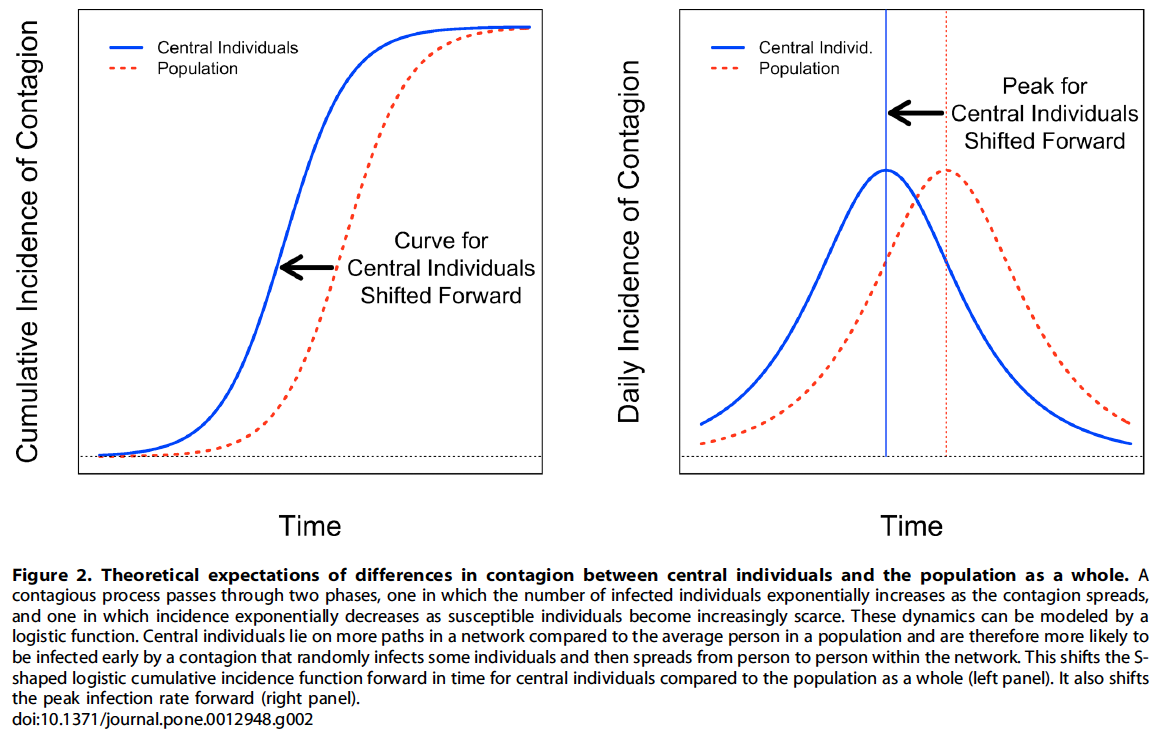
Based on this, we would expect the typical "S"—shaped infection curve for a central set of nodes, as opposed to a random sample from the population to be shifted to the left (experienced earlier in time) Fig. 2
- This shift could allow for early detection
While this may be true, the challenge lies in identifying these central nodes without mapping the entire network
Central Nodes versus Random Population
Using Friends as Proxy for Central Nodes
To solve the problem of finding central nodes, they propose a very simple solution:
Monitor the friends of those randomly selected
- These individuals are referred to as "social network sensors"
The idea here is to exploit the friendship paradox which states that, on average, our friends have more friends than we do.
"This strategy exploits an interesting property of human social networks: on average, the friends of randomly selected people possess more links (have higher degree) and are also more central (e.g., as measured by betweenness centrality) to the network than the initial, randomly selected people who named them. Therefore, we expect a set of nominated friends to get infected earlier than a set of randomly chosen individuals (who represent the population as a whole). More specifically, a random sample of individuals from a social network will have a mean degree of (the mean degree for the population); but the friends of these random individuals will have a mean degree of plus a quantity defined by the variance of the degree distribution divided by . Hence, when there is variance in degree in a population, and especially when there is high variance, the mean number of contacts for the friends will be greater (and potentially much greater) than the mean for the random sample."
Evaluating the Effectiveness of this Method
- To evaluate the effectiveness of nominated friends as social network sensors, they monitored the spread of flu at Harvard College from September 1 to December 31, 2009
Sample:
| Randomly Selected | Nominated by the Randomly Selected (Friends) | Total (Including Friends of Friends) |
|---|---|---|
| 319 | 425 | 1,789 |
In addition, as a byproduct of empaneling the foregoing group of 744 students, we wound up having information about a total of 1,789 uniquely identified Harvard College students (who either participated in the study or who were nominated as friends or as friends of friends); we used this information to draw the social network of part of the Harvard College student body
Tracked cases of influenza via:
- Formal diagnosis as well as
- Utilizing University Health Services data on the sample
"…we also collected self-reported flu symptom information from participants via email twice weekly (on Mondays and Thursdays), continuing until December 31, 2009. The students were queried about whether they had had a fever or flu symptoms since the last email contact, and there was very little missing data (47% of the subjects completed all of the biweekly surveys, and 90% missed no more than two of the surveys).
Self-report of symptoms rather than serological testing is the current standard for flu diagnosis. Similar to previous studies, students were deemed to have a case of flu (whether seasonal or the H1N1 variety) if they reported having a fever of greater than 100 F (37.8 C) and at least two of the following symptoms: sore throat; cough; stuffy or runny nose; body aches; headache; chills; or fatigue. We checked the sensitivity of our findings by using definitions of flu that required more symptoms, and our results did not change."
Important Clarification
"To be clear, we are not suggesting that a person’s precise position in the observed network, nor indeed whether he was nominated as a friend or not (and by whom), traces out the actual path by which he acquired (or did not acquire) the flu. The topological parameters we measured here, or indeed the fact that a person was deemed to be a member of the friend group, serve as proxies for the subject’s actual location within what is an essentially unobservable social network (including real friends, relatives, casual contacts, and so on) through which the flu spreads by interpersonal means. Being a ‘‘friend’’ is a marker for a person’s social network position, whatever the path of infection to this person actually is. Of course, it is likely that measured friendship networks are related to contact networks more generally: for instance, people with more friends should come into greater contact with more strangers both directly and indirectly via their friends."
Results
Numerous demographic factors were studied and shown to be not significantly related to flu diagnosis
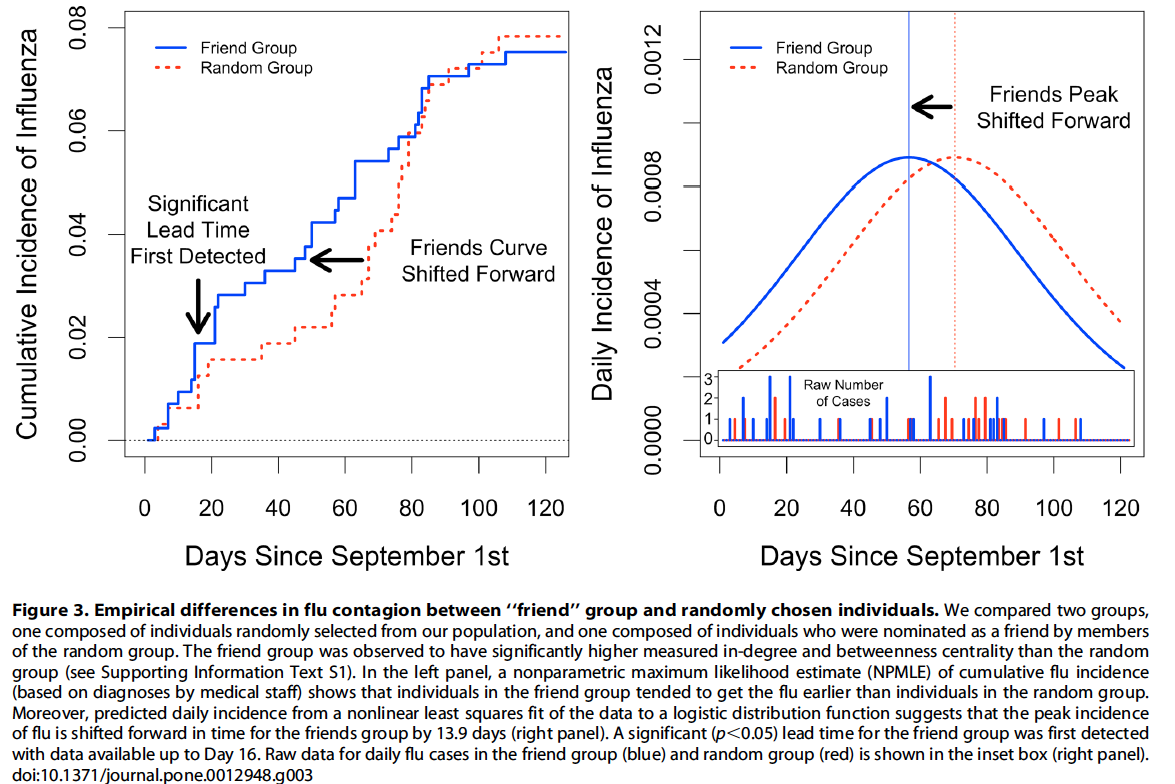
As hypothesized, the cumulative incidence curves for the friend group and the random group diverge and then converge [Figure 3]
Friends curve for flu diagnosed by medical staff is shifted 13.9 days forward in time (95% C.I. 9.9–16.6), thus providing early detection
- 60% of one standard deviation in the time-to-event in the whole sample
The results also indicate a significant but smaller shift in self-reported flu symptoms (3.2 days, 95% C.I. 2.2–4.3)
In the case of both the clinical and self-reported diagnostic standards, the estimates are robust to a number of control variables including H1N1 vaccination, seasonal flu vaccination, sex, college class, and inter-collegiate sports participation
Friend Sample versus Random Sample
Early Prediction
For flu diagnoses by medical staff, the friend group showed a significant lead time (p, 0.05) on day 16
- 46 days before the estimated peak in daily incidence in visits to the health service.
For self-reported flu symptoms, the friend group showed a significant lead time by day 39, which is 83 days prior to the estimated peak in daily incidence in self-reported symptoms.
Thus, a comparison of outcomes in the friends group and the randomly chosen group could be an effective technique for detecting outbreaks at early stages of an epidemic.
They also wondered if you could simply ask people to describe their own popularity in order to get the same predictive information. They found that this method did not work as well.
These results suggest that being nominated as a friend captures more network information (including the tendency to be central in the network) than self-reported network attributes.
Network Structure
With their data, they were able to create a small network and calculate certain features about that network such as:
- In-degree — number of times a subject was nominated as a friend
- Betweenness centrality — the number of shortest paths in the network that pass through an individual
- Coreness — the number of friends an individual has when all individuals with fewer friends are iteratively removed from the network
- Transitivity — the probability that two of one’s friends are friends with one another
They point out that this is only possible because they are sampling from a small network (Harvard) and this would likely not work if you were to do this sampling from a much larger population like a city.
- This is because the sampled people would be much less likely to known one another/be connected to one another
The results showed that, as expected, the friend group differed significantly from the random group for all these measures
- Higher in-degree (Mann Whitney U test p,0.001)
- Higher centrality (p,0.001)
- Higher k-coreness (p,0.001)
- Lower transitivity (p=0.039)
Each of these are potentially helpful measures for early prediction of spreading phenomena.
- Higher in-degree, betweenness centrality, and k—coreness should each be helpful for prediction as they are proxies for a nodes central connectedness
- The opposite is true for lower-transitivity because high transitivity means that a person has redundant network connections
They go into a bit more detail testing how each of these parameters helps for prediction in a bit more detail within the original text but I am leaving that out because it's not very important.
Couple of Discussion Points
The difference in the timing of the course of the epidemic in the friend and random groups could be exploited in at least two different ways.
- If solely the friends group were being monitored, an analyst tracking an outbreak might look for the first evidence that the incidence of the pathogen among the friends
- In a strategy that would yield more information, the analyst could track both a sample of friends and a sample of random subjects, and the harbinger of an epidemic could be taken to be when the two curves were seen to first diverge from each other
"Especially in the case of the spread of contagions other than biological pathogens, the difference between these two curves provides additional information: the adoption curve among the random sample provides evidence of secular trends in the population, whereas the difference between the two curves provides evidence of a network (inter-personal) effect, over and above the baseline force of the epidemic."
"The ability of the proposed method to detect outbreaks early, and how early it might do so, will depend on intrinsic properties of the thing that is spreading…"
Notes by Matthew R. DeVerna